Multiomics
Single Cell
Single-cell Whole Genome Sequencing

Single-cell Whole Genome Sequencing (scWGS) is a transformative technology that empowers the identification of somatic mutations present at remarkably low mosaicism levels, which is challenging for bulk sequencing.
From uncovering developmental pathways to elucidating cancer progression, scWGS offers invaluable insights into the intricacies of biology and disease.
Mirxes Genomics offers the groundbreaking scWGS service utilizing Primary Template-directed Amplification (PTA), which addresses key challenges associated with Whole Genome Amplification (WGA) and demonstrates reduced error propagation. With expert experimental and bioinformatics support from Mirxes, researchers can readily harness scWGS to achieve extraordinary single-cell clarity.
Unlock Single-cell Genomic Clarity
Partner with Us to Achieve Your Research Goals
Detect Single-cell Mosaicism
Amplify trace amounts of genomic DNA from each single cell
Exceptional WGA Performance
High genome coverage and uniformity, low error and reduced artifacts
End-to-end Service
Available from sample preparation to bioinformatics analysis
Expert Bioinformatics Support
e.g. CNV, SNV and indel analysis at single-cell level

Isolated single cell

Cell/ nuclear lysis and Whole Genome Amplification

Library construction and sequencing
Bioinformatics analysis
| Recommended sequencing depth* | Sequencing Platforms | Turnaround Time |
|---|---|---|
| 300M reads | DNBSEQ-T7 or DNBSEQ-T10 | 4-6 weeks from successful WGA QC* to FASTQ file delivery |
*This serves only as a guide. Please contact us for further discussion.
| Sample Type | Sample Source | Brief Sample Preparation Guideline* |
|---|---|---|
| Single intact cells | Fresh tissue, Fresh frozen tissue, Cell line | - Single cells/nuclei should be dispensed into 384-well plate (Lobind and Optical) - After cell dispensing, the plate should be spun down and stored at -80°C until use - Note: DAPI staining is not compatible with this assay |
| Single nuclei |
Kindly note that this serves as a brief reference only.
* Please contact us for the complete sample preparation and submission instructions.
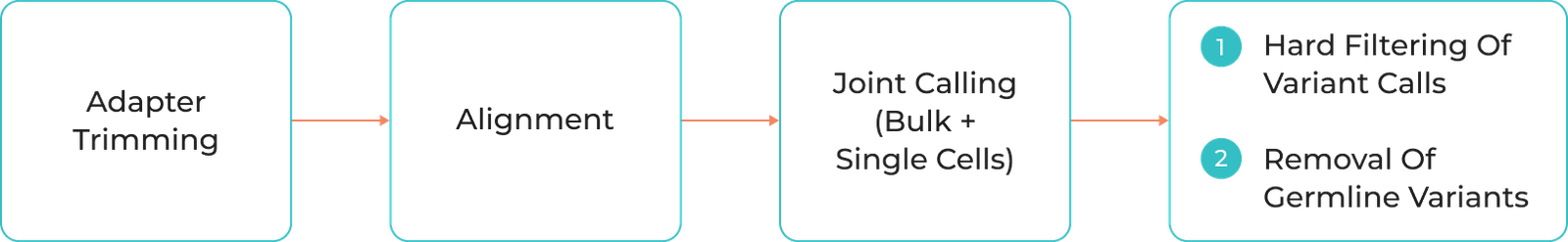
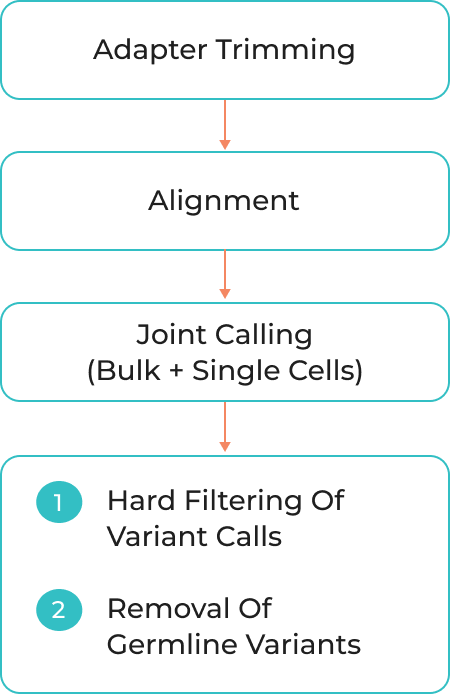
Adapter Trimming
Alignment
Joint Calling (Bulk + Single cells)
1) Hard Filtering of Variant Calls
2) Removal of germline variants
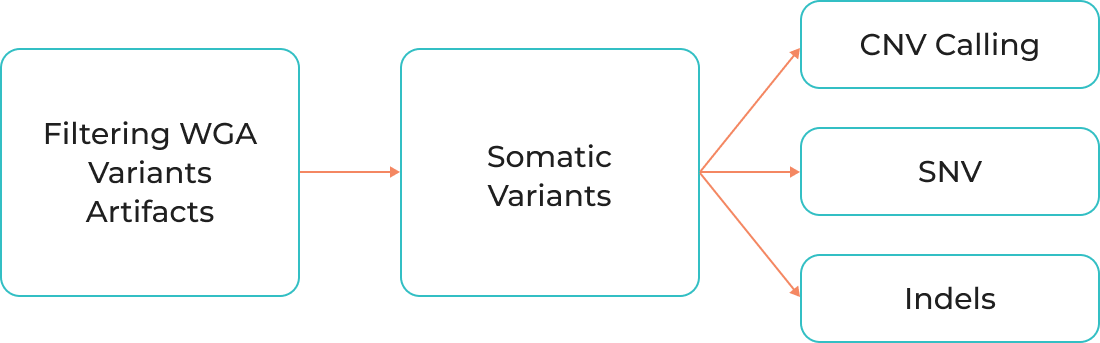
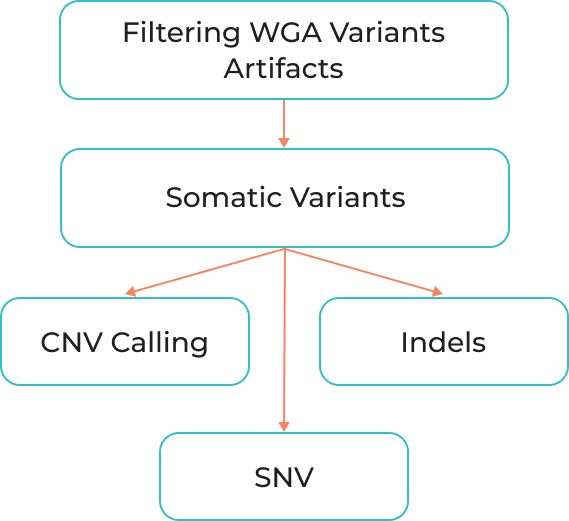
Filtering WGA variants artifacts
Somatic variants
CNV Calling
SNV
Indels
- Data Quality Control: Filtering reads with adapter or low-quality sequence data
- Alignment to reference genome using BWA
- Summary statistics
- Joint calling (Bulk and Single cells)
- Hard filtering of variant calls and removal of germline variants using paired normal bulk WGS data
- Filtering whole genome amplified variant artifacts
- Final variants: SNV, Indels and CNV
- Annotations and mutational signatures